Finding the Best Cherry Blossom Streets in Vancouver
Geospatial Analysis, Route Optimization, and Science Communication with R
2026-04-30
Motivation
- Every spring Vancouver gets buried in cherry blossoms
- The Vancouver Cherry Blossom Festival runs ~March 27 – April 17
- But which streets are actually worth visiting?
- And could you see the best of them in a single run?
The Goal
- Find all Japanese flowering cherry trees in Vancouver
- Identify the streets (and neighbourhoods) with the highest density
- Build an optimized route to visit the top streets in one trip
All code: https://github.com/chendaniely/yvr-cherry-blossoms
The Full Pipeline
data/original/ ← Vancouver Open Data (raw GeoJSON)
↓ code/01-clean/
data/final/ ← cherry_blossoms.geojson, streets.geojson
↓ code/02-street_tree/
data/processed/ ← snapped-tree-street.geojson, streets_counted.geojson
↓ results/street_results/README.qmd
data/processed/ ← top30_streets.geojson
↓ results/tsp_route/README.qmd + code/03-routes/gpx_export.R
data/routes/ ← 4 GPX filesShout Out: Kyle Walker, PhD
The project that started it all:
Kyle Walker — great R packages for spatial work:
{tidycensus},{tigris},{crsuggest}{mapgl}- every interactive map in this talk
Then I saw this post:
“New in the spopt-r #rstats package: route optimization. Solve the classic Traveling Salesman Problem for a single driver or optimize a fleet by solving the Vehicle Routing Problem.”
Find Kyle:
{spopt} - R route optimization:
Outline
Setup
- GIS concepts review
- Vancouver Open Data Portal
- The tree & street data
Analysis
- Building the initial map
- Neighbourhood-level questions
- Street-level density
- Route optimization
GIS Concepts Review
Previous R-Ladies Talks
Create Custom Webpages with R and Quarto - Grace (March 2026)
R-Ladies YVR · Meetup event
Intro to Spatial Data in R - Véronique & Johannes (2025)
R-Ladies YVR · Meetup event
If you missed those talks - this section is a quick refresher on the GIS concepts we’ll use today.
What is GIS?
Geographic Information System - tools for capturing, storing, analyzing spatial data
Vector data (what we use today)
- Points - individual trees 🌸
- Lines - street segments 🛣️
- Polygons - neighbourhoods 🗺️
Raster data (not today)
- Grids of pixels
- Satellite imagery, elevation
{terra},{stars}
Coordinate Reference Systems
Every spatial dataset needs a CRS - defines how coordinates map to Earth’s surface
Warning
Mix up your CRS and you’ll get wrong results with no error message.
Coordinate Reference Systems
Reading layer `public-trees' from data source
`/Users/dan/git/hub/rladies-2026-04-30-blossoms/submodule/yvr-cherry-blossoms/data/original/public-trees.geojson'
using driver `GeoJSON'
replacing null geometries with empty geometries
Simple feature collection with 186342 features and 11 fields (with 1 geometry empty)
Geometry type: POINT
Dimension: XY
Bounding box: xmin: -123.2367 ymin: 49.2002 xmax: -123.0233 ymax: 49.30998
Geodetic CRS: WGS 84Coordinate Reference System:
User input: WGS 84
wkt:
GEOGCRS["WGS 84",
DATUM["World Geodetic System 1984",
ELLIPSOID["WGS 84",6378137,298.257223563,
LENGTHUNIT["metre",1]]],
PRIMEM["Greenwich",0,
ANGLEUNIT["degree",0.0174532925199433]],
CS[ellipsoidal,2],
AXIS["geodetic latitude (Lat)",north,
ORDER[1],
ANGLEUNIT["degree",0.0174532925199433]],
AXIS["geodetic longitude (Lon)",east,
ORDER[2],
ANGLEUNIT["degree",0.0174532925199433]],
ID["EPSG",4326]]CRS Alphanumeric Soup
EPSG:4326 = WGS84
- EPSG - a registry of named coordinate systems (like a phone book for CRS)
- 4326 - the ID for WGS84 in that registry
- WGS84 - the actual name: World Geodetic System 1984
- What you get: lat/lon in degrees, used by GPS, Google Maps, GeoJSON
EPSG:32610 = UTM Zone 10N
- 32610 - EPSG ID for UTM Zone 10N (covers BC/Pacific Northwest)
- UTM - Universal Transverse Mercator, a family of projected systems
- Zone 10N - the specific slice of the globe (Vancouver falls here)
- What you get: X/Y in metres - essential for distance calculations
Tip
Rule of thumb: use 4326 for display/storage, 32610 for anything involving distance or area in BC.
Key {sf} Operations We’ll Use
| Function | What it does |
|---|---|
st_read() |
Load geojson / shapefile / etc |
st_transform() |
Reproject to a new CRS |
st_nearest_feature() |
Snap each point to closest line |
st_length() |
Length of line in CRS units |
st_join() |
Spatial join (like SQL JOIN but spatial) |
st_buffer() |
Buffer geometries by a distance |
st_union() |
Merge geometries together |
Key R Packages
library(sf) # spatial data I/O and operations
library(dplyr) # data manipulation
library(mapgl) # interactive maps (MapLibre GL)
library(sfnetworks) # street/graph networks
library(igraph) # graph algorithms (TSP cost matrix)
library(spopt) # route optimization (TSP solver)
library(osmdata) # OpenStreetMap data queriesKey R Packages
Vancouver Open Data Portal
opendata.vancouver.ca
https://opendata.vancouver.ca/pages/home/
The City of Vancouver publishes hundreds of datasets under open licenses:
- Trees, parks, bike lanes, zoning, permits, 3D buildings…
We’ll use 5 datasets today:
| Dataset | Format | What it contains |
|---|---|---|
public-trees |
GeoJSON | All ~150k city trees |
public-streets |
GeoJSON | City-maintained streets |
non-city-streets |
GeoJSON | Private/strata streets |
lanes |
GeoJSON | Lane segments |
local-area-boundary |
GeoJSON | 22 neighbourhood polygons |
Reading the Data
Reading the Data
Rows: 186,342
Columns: 12
$ asset_id <int> 270046, 270058, 270059, 270079, 270081, 270085, 270090, …
$ address <chr> "6110 ST. CLAIR PLACE", "3444 ST. CATHERINES ST", "3104 …
$ common_name <chr> "CHERRY, PLUM OR PEACH SPECIES", "JAPANESE SNOWBELL", "W…
$ genus_name <chr> "PRUNUS", "STYRAX", "TSUGA", "FRAXINUS", "PARROTIA", "FR…
$ species_name <chr> "SPECIES", "JAPONICUS", "HETEROPHYLLA", "ORNUS", "PERSIC…
$ cultivar_name <chr> "NONE", "NONE", "NONE", "ARIE PETERS", "PERSIAN SPIRE", …
$ height_m <dbl> 13.7, 4.6, 19.8, 4.6, 4.6, 5.0, 4.6, 4.6, 4.6, 4.6, 4.6,…
$ diameter_cm <dbl> 88.9, 7.6, 66.0, 5.1, 7.6, 8.0, 7.6, 7.6, 7.6, 7.6, 7.6,…
$ date_planted <date> NA, 2021-12-15, NA, 2023-03-09, 2021-03-25, 2022-12-16,…
$ geom <chr> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, …
$ geo_point_2d <chr> "{ \"lon\": -123.19037399947032, \"lat\": 49.23186499926…
$ geometry <POINT [°]> POINT (-123.1904 49.23186), POINT (-123.0856 49.25…Reading the Data
Reading layer `public-trees' from data source
`/Users/dan/git/hub/rladies-2026-04-30-blossoms/submodule/yvr-cherry-blossoms/data/original/public-trees.geojson'
using driver `GeoJSON'
replacing null geometries with empty geometries
Simple feature collection with 186342 features and 11 fields (with 1 geometry empty)
Geometry type: POINT
Dimension: XY
Bounding box: xmin: -123.2367 ymin: 49.2002 xmax: -123.0233 ymax: 49.30998
Geodetic CRS: WGS 84Reading layer `public-streets' from data source
`/Users/dan/git/hub/rladies-2026-04-30-blossoms/submodule/yvr-cherry-blossoms/data/original/public-streets.geojson'
using driver `GeoJSON'
Simple feature collection with 17067 features and 3 fields
Geometry type: MULTILINESTRING
Dimension: XY
Bounding box: xmin: -123.2248 ymin: 49.20032 xmax: -123.0233 ymax: 49.2945
Geodetic CRS: WGS 84Reading layer `non-city-streets' from data source
`/Users/dan/git/hub/rladies-2026-04-30-blossoms/submodule/yvr-cherry-blossoms/data/original/non-city-streets.geojson'
using driver `GeoJSON'
Simple feature collection with 952 features and 4 fields
Geometry type: LINESTRING
Dimension: XY
Bounding box: xmin: -123.2299 ymin: 49.19895 xmax: -123.0124 ymax: 49.31404
Geodetic CRS: WGS 84Reading layer `lanes' from data source
`/Users/dan/git/hub/rladies-2026-04-30-blossoms/submodule/yvr-cherry-blossoms/data/original/lanes.geojson'
using driver `GeoJSON'
Simple feature collection with 7851 features and 3 fields
Geometry type: LINESTRING
Dimension: XY
Bounding box: xmin: -123.2223 ymin: 49.20096 xmax: -123.0233 ymax: 49.29374
Geodetic CRS: WGS 84Reading layer `local-area-boundary' from data source
`/Users/dan/git/hub/rladies-2026-04-30-blossoms/submodule/yvr-cherry-blossoms/data/original/local-area-boundary.geojson'
using driver `GeoJSON'
Simple feature collection with 22 features and 2 fields
Geometry type: POLYGON
Dimension: XY
Bounding box: xmin: -123.2248 ymin: 49.19894 xmax: -123.0232 ymax: 49.29581
Geodetic CRS: WGS 84The Trees Data
Which Trees Are Cherry Blossoms?
150k+ trees in the dataset - most are not cherry blossoms.
Note
I had no idea there were so many different kinds of cherry blossom trees!
Strategy: filter common_name for trees that mention “cherry” AND (“japanese” OR “flower”)
Which Trees Are Cherry Blossoms?
[1] "KWANZAN FLOWERING CHERRY" "AKEBONO FLOWERING CHERRY"
[3] "SARGENT FLOWERING CHERRY" "JAPANESE FLOWERING CHERRY"
[5] "UKON JAPANESE CHERRY" "AMANOGAWA JAPANESE CHERRY"
[7] "WEEPING JAPANESE CHERRY" NA
[9] "OJOCHIN JAPANESE CHERRY" "SHOGETSU JAPANESE CHERRY"
[11] "DOUBLE WEEPING JAPANESE CHERRY"Filtering the Trees
10 species found:
JAPANESE FLOWERING CHERRYAKEBONO FLOWERING CHERRYKWANZAN FLOWERING CHERRY- … and more
Filtering the Trees
[1] 14992Data Quality: A Frustration
Mature trees are more reliable bloomers. Filter by age?
n has_date pct_missing
3521 979 72.2Warning
72% of trees have no planting date - age filtering is off the table.
Data Quality: A Frustration
Simple feature collection with 1 feature and 3 fields
Geometry type: MULTIPOINT
Dimension: XY
Bounding box: xmin: -123.2319 ymin: 49.20477 xmax: -123.0236 ymax: 49.30261
Geodetic CRS: WGS 84
n has_date pct_missing geometry
1 14992 4113 72.56537 MULTIPOINT ((-123.0256 49.2...Falling Back to Size Proxies
Height and diameter are fully recorded - use them as size stand-ins:
# Thresholds (based on domain knowledge of Japanese flowering cherries)
min_height_m <- 2.5 # metres
min_diameter_cm <- 7 # cm trunk diameter
# NOTE: in the final analysis, even filtering by these thresholds
# changes the result very little - most trees pass. We keep all trees.
blooming_trees <- cherry_blossomsFalling Back to Size Proxies
Falling Back to Size Proxies
Simple feature collection with 1 feature and 2 fields
Geometry type: MULTIPOINT
Dimension: XY
Bounding box: xmin: -123.2319 ymin: 49.20477 xmax: -123.0236 ymax: 49.30261
Geodetic CRS: WGS 84
pct_height pct_diameter geometry
1 100 100 MULTIPOINT ((-123.0256 49.2...The Street Data
Three sources - combine them into one dataset:
streets_pub <- st_read(
"submodule/yvr-cherry-blossoms/data/original/public-streets.geojson",
quiet = TRUE
) |> st_transform(4326) |>
mutate(street_type = "Public", street_label = hblock)
streets_non <- st_read(
"submodule/yvr-cherry-blossoms/data/original/non-city-streets.geojson",
quiet = TRUE
) |> st_transform(4326) |>
mutate(
street_type = "Non-city",
street_label = if_else(
!is.na(from_hblk) & from_hblk != "",
paste(from_hblk, streetname), streetname
)
)
streets_lan <- st_read(
"submodule/yvr-cherry-blossoms/data/original/lanes.geojson",
quiet = TRUE
) |> st_transform(4326) |>
mutate(street_type = "Lane",
street_label = paste(from_hundred_block, std_street))
streets <- bind_rows(streets_pub, streets_non, streets_lan)
nrow(streets)Note: each street is stored as individual block-level segments
The Street Data
[1] 25870Building the Initial Map
First Map: Where Are the Trees?
Start simple - plot all cherry blossom trees with {mapgl}
First Map: Where Are the Trees?
Adding Neighbourhood Boundaries
maplibre(
style = openfreemap_style("bright"),
bounds = neighbourhoods
) |>
add_line_layer(
id = "neighbourhood-borders",
source = neighbourhoods,
line_color = "#d97706",
line_width = 1.5,
line_opacity = 0.8
) |>
add_circle_layer(
id = "trees",
source = cherry_blossoms,
circle_radius = 3,
circle_color = "#e879a0",
circle_opacity = 0.6
)Adding Neighbourhood Boundaries
All Cherry Blossoms by Species
tree_variety_names <- sort(unique(cherry_blossoms$common_name))
tree_variety_colors <- colorRampPalette(
c("#e879a0", "#a855f7", "#3b82f6", "#10b981", "#f59e0b")
)(length(tree_variety_names))
maplibre(
style = openfreemap_style("bright"),
bounds = cherry_blossoms
) |>
add_circle_layer(
id = "trees",
source = cherry_blossoms,
circle_color = match_expr(
column = "common_name",
values = tree_variety_names,
stops = tree_variety_colors
),
circle_opacity = 0.7,
tooltip = concat("<b>", get_column("common_name"), "</b>")
) |>
add_categorical_legend(
legend_title = "Tree variety",
values = tree_variety_names,
colors = tree_variety_colors,
patch_shape = "circle"
)All Cherry Blossoms by Species
Asking Questions
Which Neighbourhood Has Most?
We need to spatially join trees to neighbourhoods, then count:
tree_counts <- st_join(
cherry_blossoms,
neighbourhoods |> select(neighbourhood = name),
join = st_within # tree must fall INSIDE neighbourhood polygon
) |>
st_drop_geometry() |>
count(neighbourhood, name = "tree_count")
neighbourhoods <- neighbourhoods |>
left_join(tree_counts, by = c("name" = "neighbourhood")) |>
mutate(tree_count = replace_na(tree_count, 0))Which Neighbourhood Has Most?
Raw Count vs Density
Raw count favours large neighbourhoods. Normalize by area:
Top by raw count
Kitsilano, Hastings-Sunrise, Sunset…
Top by density
Mount Pleasant (!)
Raw Count vs Density
The Neighbourhood Choropleth
maplibre(style = openfreemap_style("bright"), bounds = neighbourhoods) |>
add_fill_layer(
id = "neighbourhood-density",
source = neighbourhoods,
fill_color = interpolate(
column = "trees_per_km2",
values = c(0, median(neighbourhoods$trees_per_km2),
max(neighbourhoods$trees_per_km2)),
stops = c("#f7fcf0", "#41ab5d", "#00441b")
),
fill_opacity = 0.75,
tooltip = "tooltip_text"
)The Neighbourhood Choropleth
Surprising Finding #1
Mount Pleasant = highest density neighbourhood
David Lam Park (Downtown) = where the Festival is held
Why the paradox?
- Downtown is geographically massive
- Trees are concentrated in one small park
- Density = trees ÷ area → big area = low density
- But Mount Pleasant has trees spread across many residential streets
Which Streets Are Best?
Neighbourhoods give the overview - the actual experience is at street level.
Where do I take my mom?
Where can folks take photos?
Where are the hidden local gems?
- Assign each tree to its nearest street segment
- Count trees per segment
- Normalize by length → trees per 100m
- Rank and map
Street-Level Analysis
Example: Snapping - Toy Setup
Build a mini version with 4 streets and 9 trees:
street_colors <- c(
"Oak St" = "#e76f51",
"Elm Ave" = "#2a9d8f",
"Cedar Rd" = "#457b9d",
"Maple Blvd" = "#9b5de5"
)
streets_toy <- st_sf(
id = 1:4,
name = c("Oak St", "Elm Ave", "Cedar Rd", "Maple Blvd"),
geometry = st_sfc(
st_linestring(rbind(c(0, 0), c(10, 0))), # horizontal
st_linestring(rbind(c(0, 8), c(10, 8))), # horizontal (upper)
st_linestring(rbind(c(5, -1), c(5, 10))), # vertical
st_linestring(rbind(c(0, 5), c(4, 10))), # diagonal
crs = 4326
)
)
trees_toy <- st_sf(
id = 1:9,
geometry = st_sfc(
st_point(c(1, -1.2)), st_point(c(3, 1.0)), st_point(c(7, -0.8)),
st_point(c(2, 4.0)), st_point(c(8, 4.5)), st_point(c(3, 7.0)),
st_point(c(6, 9.2)), st_point(c(9, 6.5)), st_point(c(4.5, 3.5)),
crs = 4326
)
)Example: Snapping - Toy Setup
Example: Before Snapping
ggplot() +
geom_sf(data = streets_toy, aes(color = name), linewidth = 2.5) +
geom_sf_text(
data = streets_toy,
aes(label = name, color = name),
size = 3.5, show.legend = FALSE
) +
geom_sf(data = trees_toy, size = 4, shape = 16, color = "grey30") +
scale_color_manual(values = street_colors, name = "Street") +
labs(
title = "Streets and trees (before snapping)",
subtitle = "Which street does each point belong to?"
) +
theme_minimal() +
theme(legend.position = "bottom")Example: Before Snapping
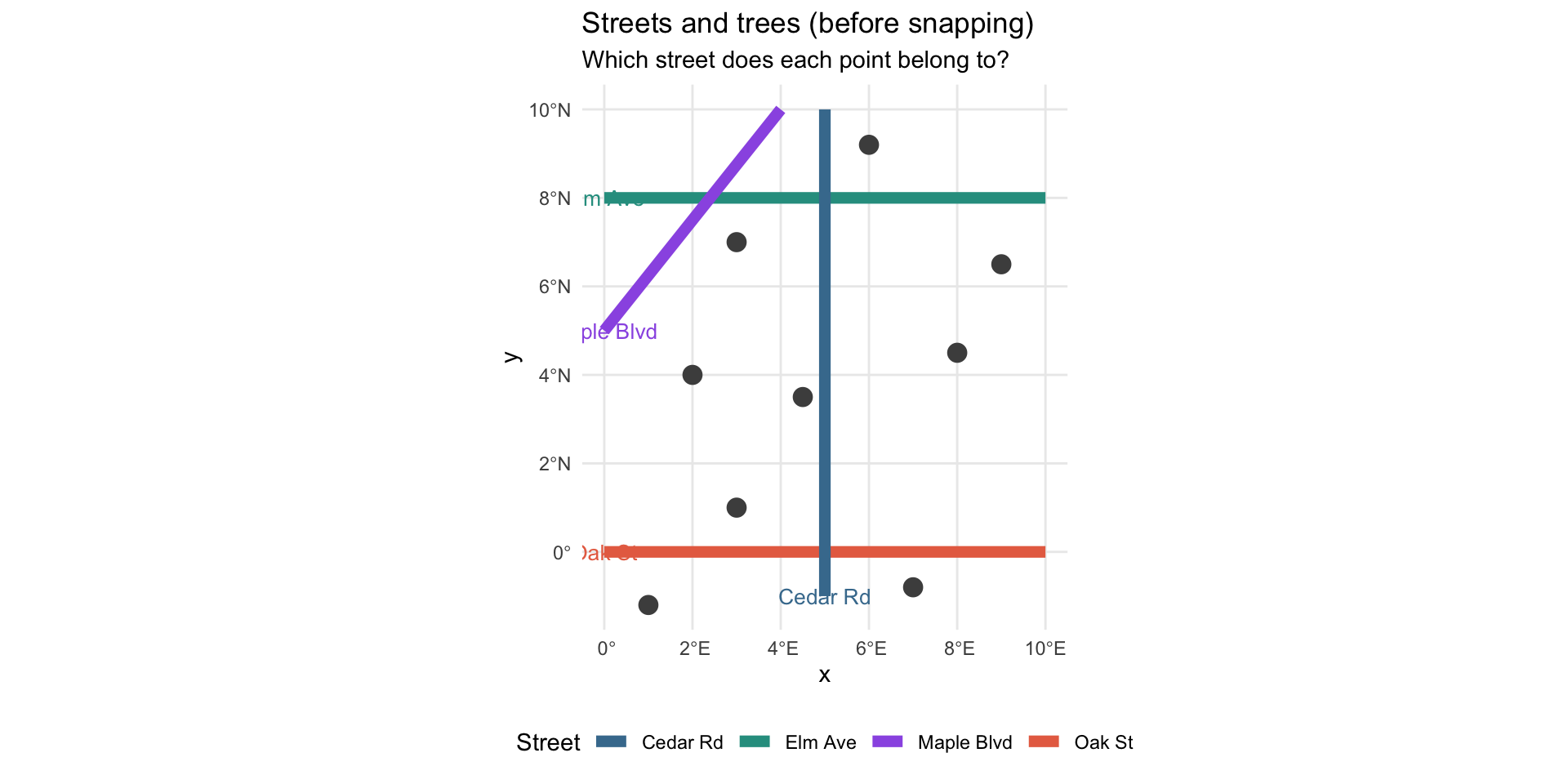
Example: The Snap Operation
# One call assigns every tree to its nearest street
nearest_idx <- st_nearest_feature(trees_toy, streets_toy)
trees_toy <- trees_toy |>
mutate(
nearest_street_id = nearest_idx,
nearest_street_name = streets_toy$name[nearest_idx]
)
tree_counts_toy <- trees_toy |>
st_drop_geometry() |>
count(nearest_street_id, nearest_street_name, name = "tree_count")
print(tree_counts_toy)Example: The Snap Operation
nearest_street_id nearest_street_name tree_count
1 1 Oak St 3
2 2 Elm Ave 2
3 3 Cedar Rd 3
4 4 Maple Blvd 1Example: After Snapping
snap_lines <- trees_toy |>
mutate(
geometry = st_sfc(
map2(
geometry,
streets_toy$geometry[nearest_street_id],
~ st_nearest_points(st_sfc(.x, crs = 4326), st_sfc(.y, crs = 4326))[[1]]
),
crs = 4326
)
)
ggplot() +
geom_sf(data = streets_toy, aes(color = name), linewidth = 2.5) +
geom_sf(data = snap_lines, color = "grey55", linetype = "dashed", linewidth = 0.6) +
geom_sf(data = trees_toy, aes(color = nearest_street_name), size = 4, shape = 16) +
geom_sf_label(
data = left_join(streets_toy, tree_counts_toy, by = c("id" = "nearest_street_id")),
aes(label = paste0(name, "\n(", tree_count, " trees)"), color = name),
size = 3.5, nudge_y = -1.2, show.legend = FALSE
) +
scale_color_manual(values = street_colors, name = "Nearest street") +
labs(
title = "Snapping points to nearest line segment",
subtitle = "Dashed lines show each point's assignment to its closest street"
) +
theme_minimal() +
theme(legend.position = "bottom")Example: After Snapping
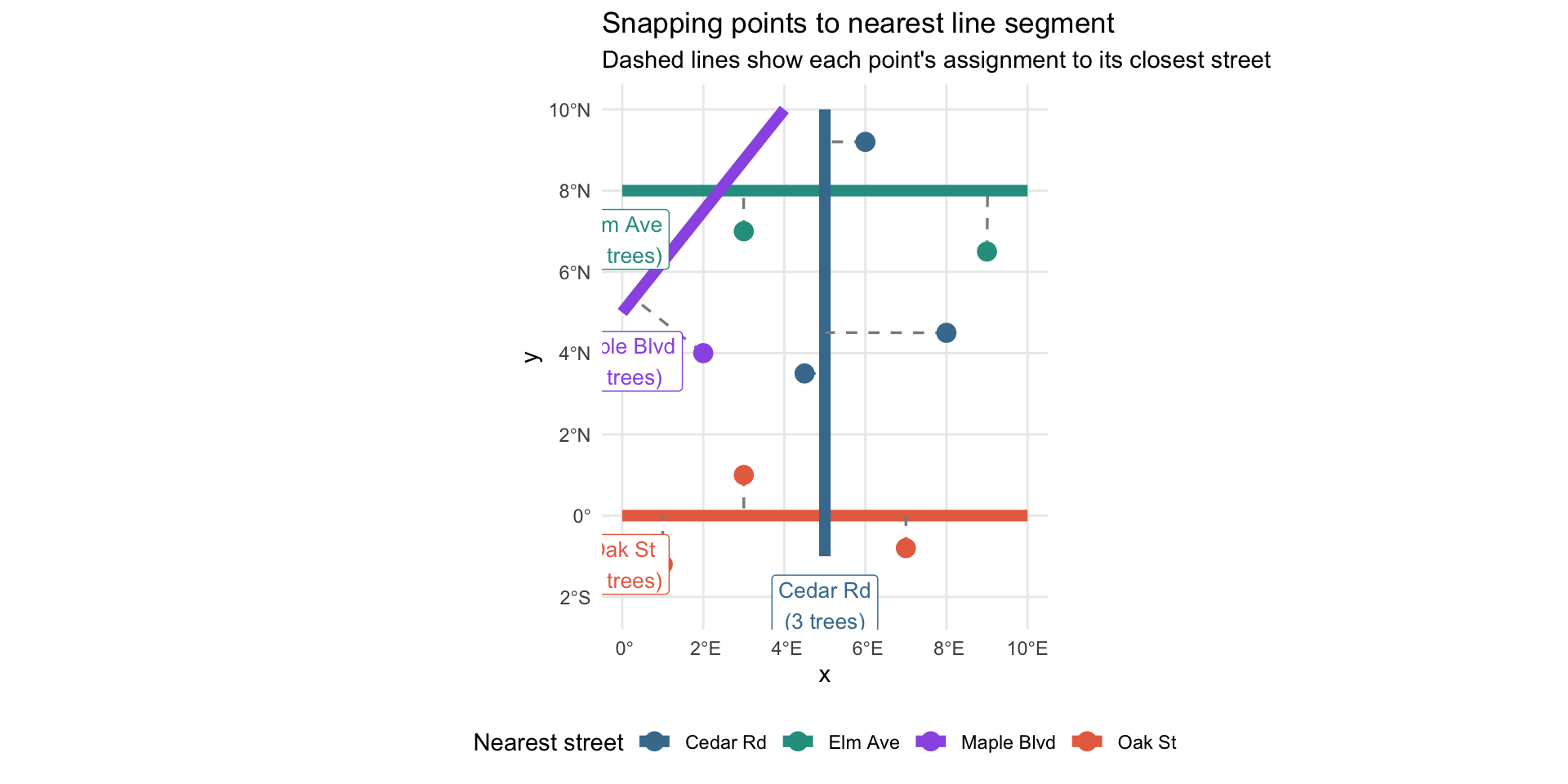
Snapping Trees to Streets
st_nearest_feature() returns the index of the nearest feature - one call for all 3,500 trees:
Note
Uses a spatial index under the hood - 3,500 trees × 50,000+ segments runs in seconds.
Snapping Trees to Streets
Counting Trees per Segment
tree_counts <- snapped |>
st_drop_geometry() |>
count(street_id, street_label, street_type,
name = "tree_count")
streets_joined <- streets |>
mutate(street_id = row_number()) |>
left_join(tree_counts, by = "street_id")
# drop duplicate .y columns, rename .x → clean names, then compute length/density
streets_counted <- streets_joined |>
select(-street_label.y, -street_type.y) |>
rename(street_label = street_label.x, street_type = street_type.x) |>
mutate(
tree_count = replace_na(tree_count, 0),
length_m = as.numeric(st_length(geometry)),
density = tree_count / length_m * 100 # trees per 100m
)Counting Trees per Segment
Data Integrity Check
After the join, verify labels are consistent across segments:
label_mismatches <- streets_joined |>
st_drop_geometry() |>
filter(!is.na(street_label.y)) |>
filter(street_label.x != street_label.y)
if (nrow(label_mismatches) > 0) {
warning(sprintf(
"%d segment(s) have mismatched labels - re-run clean scripts",
nrow(label_mismatches)
))
} else {
message("street_label check passed ✓")
}Tip
Always validate spatial joins - silent mismatches are common.
Data Integrity Check
Example: Block-Level Segments
One “street” in the data is many separate block segments - pick one to see:
Note
Each colour is a separate row in the data - that’s why we need to aggregate before ranking streets.
Example: Block-Level Segments
Aggregating to Street Level
Each street has multiple block segments. Union them to see the whole street:
# summary table (no geometry - used for ranking)
street_summary <- streets_counted |>
st_drop_geometry() |>
filter(tree_count > 0) |>
group_by(street_label, street_type) |>
summarise(
total_trees = sum(tree_count),
total_length = sum(length_m),
n_segments = n(),
.groups = "drop"
) |>
mutate(density = total_trees / total_length * 100)
# spatial version (geometry needed for mapping)
streets_sf <- streets_counted |>
filter(tree_count > 0) |>
group_by(street_label, street_type) |>
summarise(
total_trees = sum(tree_count),
total_length = sum(length_m),
density = sum(tree_count) / sum(length_m) * 100,
geometry = st_union(geometry),
.groups = "drop"
)Aggregating to Street Level
Top Streets
# Top by raw count
top_by_count <- street_summary |>
slice_max(total_trees, n = 20)
# Top by density - require ≥5 trees to suppress short stubs
# (a single tree on a 5m dead-end = 2000 trees/100m, meaningless)
top_by_density <- street_summary |>
filter(total_trees >= 5) |>
slice_max(density, n = 30)Important
Without total_trees >= 5, a single tree on a 5m dead-end alley tops the rankings at 2,000 trees/100m.
Top Streets
The Top 30 Streets Map
maplibre(style = openfreemap_style("bright"), bounds = top30_stops) |>
add_line_layer(
id = "streets-density",
source = streets_dense,
line_color = interpolate(
column = "density",
values = c(0, median(streets_dense$density),
max(streets_dense$density)),
stops = c("#edf8e9", "#74c476", "#00441b")
),
line_width = interpolate(
column = "density",
values = c(0, max(streets_dense$density)),
stops = c(1.5, 6)
),
popup = concat(
"<b>", get_column("street_label"), "</b><br/>",
"Trees / 100m: ", number_format(get_column("density"), 1)
)
)The Top 30 Streets Map
Surprising Finding #2
Mount Pleasant has the highest neighbourhood density
but none of the top 30 densest streets
Why? Mount Pleasant trees are spread across many streets - great for wandering, but no single street is exceptionally dense.
The top streets cluster in West End / Yaletown, Kerrisdale, and East Vancouver.
The Top 10 Streets
| # | Street | Trees / 100m | Total Trees |
|---|---|---|---|
| 1 | 1400 HOMER ST | 27.5 | 53 |
| 2 | 100 DRAKE ST | 22.0 | 25 |
| 3 | 600 GILFORD ST | 22.4 | 13 |
| 4 | 600 CHILCO ST | 24.6 | 16 |
| 5 | 4500 BALACLAVA ST | 22.7 | 12 |
| 6 | 2400 W 41ST AV | 24.4 | 25 |
| 7 | 400 W 33RD AV | 20.5 | 18 |
| 8 | 200 W 37TH AV | 21.6 | 21 |
| 9 | 800 SALSBURY DRIVE | 22.0 | 11 |
| 10 | 3100 RUPERT ST | 19.3 | 18 |
Route Optimization
Travelling Salesperson Problem
Given N stops, find the shortest route visiting each exactly once.
- NP-hard exactly - heuristics for anything practical
{spopt}by Kyle Walker: nearest-neighbour seed + 2-opt/or-opt improvement- Key input: a cost matrix of distances or travel times between every pair of stops
10 stops → 10! / 2 = 1,814,400 possible routes

Running the TSP
coords <- st_coordinates(top10_stops)
cost_matrix <- as.matrix(dist(coords)) # Euclidean - fast, good for prototyping
result <- route_tsp(
top10_stops,
start = 1, # Homer St (#1 ranked)
end = NULL, # open route - no return to start
cost_matrix = cost_matrix
)
meta <- attr(result, "spopt")
cat(sprintf("Improvement over nearest-neighbour: %.1f%%\n", meta$improvement_pct))Running the TSP
Improvement over nearest-neighbour: 16.8%Simple feature collection with 10 features and 2 fields
Geometry type: POINT
Dimension: XY
Bounding box: xmin: -123.1757 ymin: 49.23468 xmax: -123.0338 ymax: 49.2943
Geodetic CRS: WGS 84
street_label .visit_order geometry
1 1400 HOMER ST 1 POINT (-123.1262 49.27227)
2 100 DRAKE ST 2 POINT (-123.1231 49.27308)
3 600 GILFORD ST 3 POINT (-123.135 49.2938)
4 600 CHILCO ST 4 POINT (-123.137 49.2943)
5 4500 BALACLAVA ST 5 POINT (-123.1757 49.24542)
6 2400 W 41ST AV 6 POINT (-123.162 49.23468)
7 400 W 33RD AV 7 POINT (-123.1163 49.24101)
8 200 W 37TH AV 8 POINT (-123.1112 49.23711)
9 800 SALSBURY DRIVE 9 POINT (-123.0684 49.27731)
10 3100 RUPERT ST 10 POINT (-123.0338 49.25599)Routing Geometry Was Off
route_tsp() returns the stop order - not the road paths between stops.
Drawing the actual route requires geometry connecting each stop in sequence.
What I tried
Build a road network from Vancouver street segments + sfnetworks, compute shortest paths, connect the dots
What happened
Routes didn’t follow roads - lines cut through blocks, missed turns, looked wrong
Why? Street Data ≠ Routing Data
- Vancouver open data: block-level records - built for spatial joins, not routing
- Tried full OpenStreetMap data - still had issues
- Gaps in network topology, or geometry too messy for a DIY routing engine
The lesson: routing is a solved problem. Don’t rebuild it from scratch.
OSRM to the Rescue
TSP gives the order - OSRM gives actual road geometry:
library(httr)
osrm_route <- function(stops_sf, profile = "foot") {
coords <- st_coordinates(stops_sf)
coord_str <- paste(
sprintf("%.6f,%.6f", coords[, 1], coords[, 2]),
collapse = ";"
)
url <- sprintf(
"https://router.project-osrm.org/route/v1/%s/%s?overview=full&geometries=geojson",
profile, coord_str
)
content(GET(url))$routes[[1]]
}Profiles: "foot" (sidewalks + paths) · "bike" (cycle lanes preferred)
OSRM to the Rescue
GPX Export
Make the route portable - Garmin, Strava, Komoot, AllTrails:
library(xml2)
write_gpx <- function(stops_sf, track_coords, name, path) {
doc <- xml_new_root("gpx", version = "1.1",
creator = "yvr-cherry-blossoms",
xmlns = "http://www.topografix.com/GPX/1/1")
pts <- st_coordinates(stops_sf)
for (i in seq_len(nrow(stops_sf))) {
wpt <- xml_add_child(doc, "wpt",
lat = sprintf("%.6f", pts[i, 2]),
lon = sprintf("%.6f", pts[i, 1]))
xml_add_child(wpt, "name",
sprintf("Stop %d - %s", i, stops_sf$street_label[i]))
}
# ... track points from OSRM geometry added here
write_xml(doc, path)
}GPX Export
Cherry Blossom Marathon Route
Top 10 stops · 23.4 km (14.5 miles) straight-line · ~35 km road-following

Top 10 cherry blossom marathon route
For Cyclists: The Grand Tour
Top 30 stops · 34.7 km (21.6 miles) straight-line
Google Maps limits routes to 10 stops - split into 3 legs:
bike_stops <- tsp_result_30 |>
filter(!is.na(.visit_order)) |>
arrange(.visit_order) |>
mutate(leg = ceiling(row_number() / 10)) # legs 1, 2, 3
# Generate Google Maps URL per leg
google_maps_url <- function(stops_sf) {
stops_sf$google_address |> # "HOMER ST, Vancouver, BC"
URLencode(reserved = TRUE) |>
paste(collapse = "/") |>
paste0("https://www.google.com/maps/dir/", x = _)
}
urls <- lapply(1:3, \(i) google_maps_url(filter(bike_stops, leg == i)))
urls[[1]]For Cyclists: The Grand Tour
[1] "https://www.google.com/maps/dir/1400%20HOMER%20ST%2C%20Vancouver%2C%20BC/100%20DRAKE%20ST%2C%20Vancouver%2C%20BC/600%20CHILCO%20ST%2C%20Vancouver%2C%20BC/600%20GILFORD%20ST%2C%20Vancouver%2C%20BC/1000%20MELVILLE%20ST%2C%20Vancouver%2C%20BC/800%20SALSBURY%20DRIVE%2C%20Vancouver%2C%20BC/2500%20OXFORD%20ST%2C%20Vancouver%2C%20BC/3000%20E%202ND%20AV%2C%20Vancouver%2C%20BC/3000%20E%207TH%20AV%2C%20Vancouver%2C%20BC/2600%20E%2022ND%20AV%2C%20Vancouver%2C%20BC"Reflections
I Actually Ran It
Still in Bud: March 30th
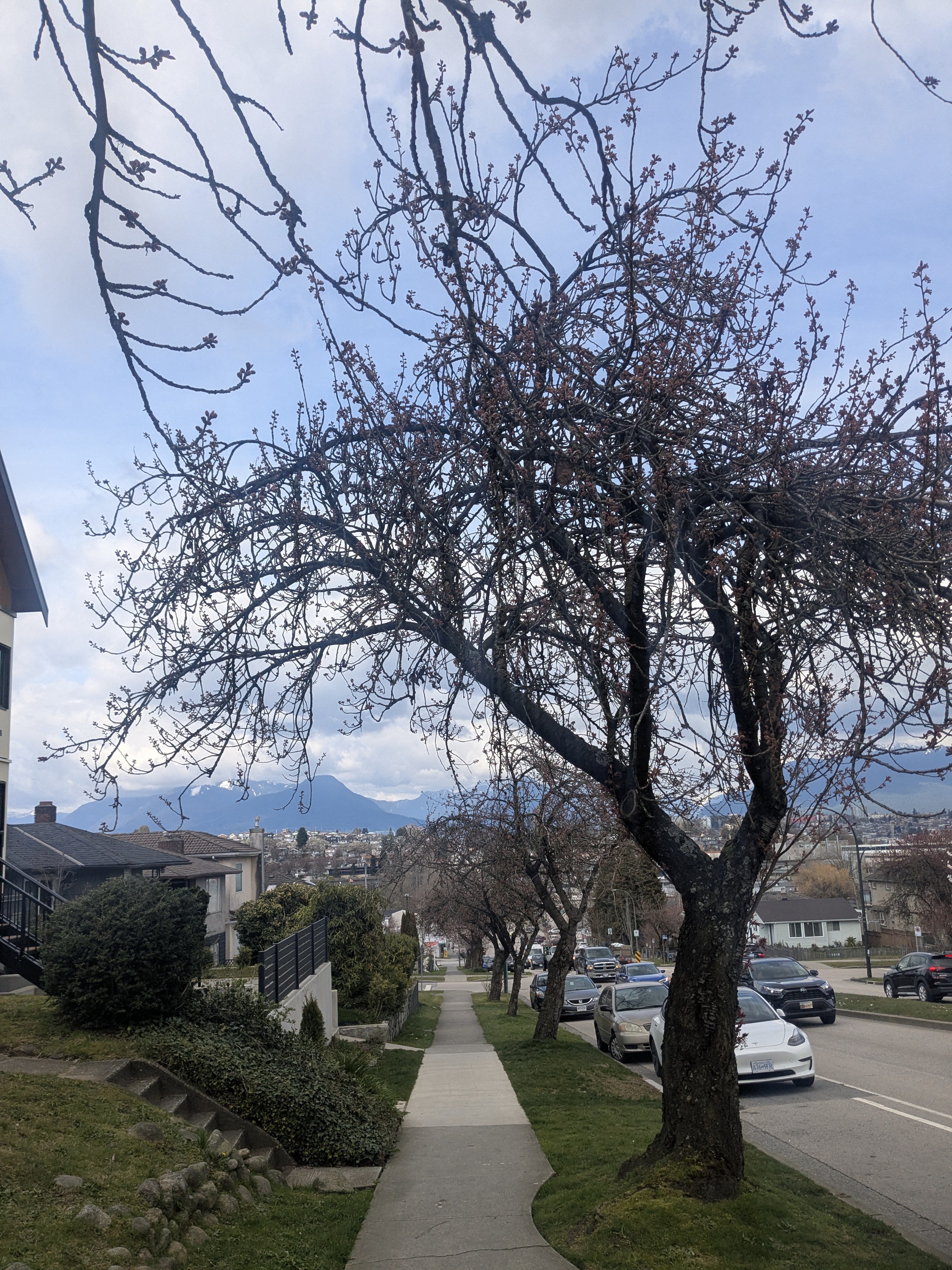


Peak Bloom: April 18th
Rupert St - with my mom, three weeks later:
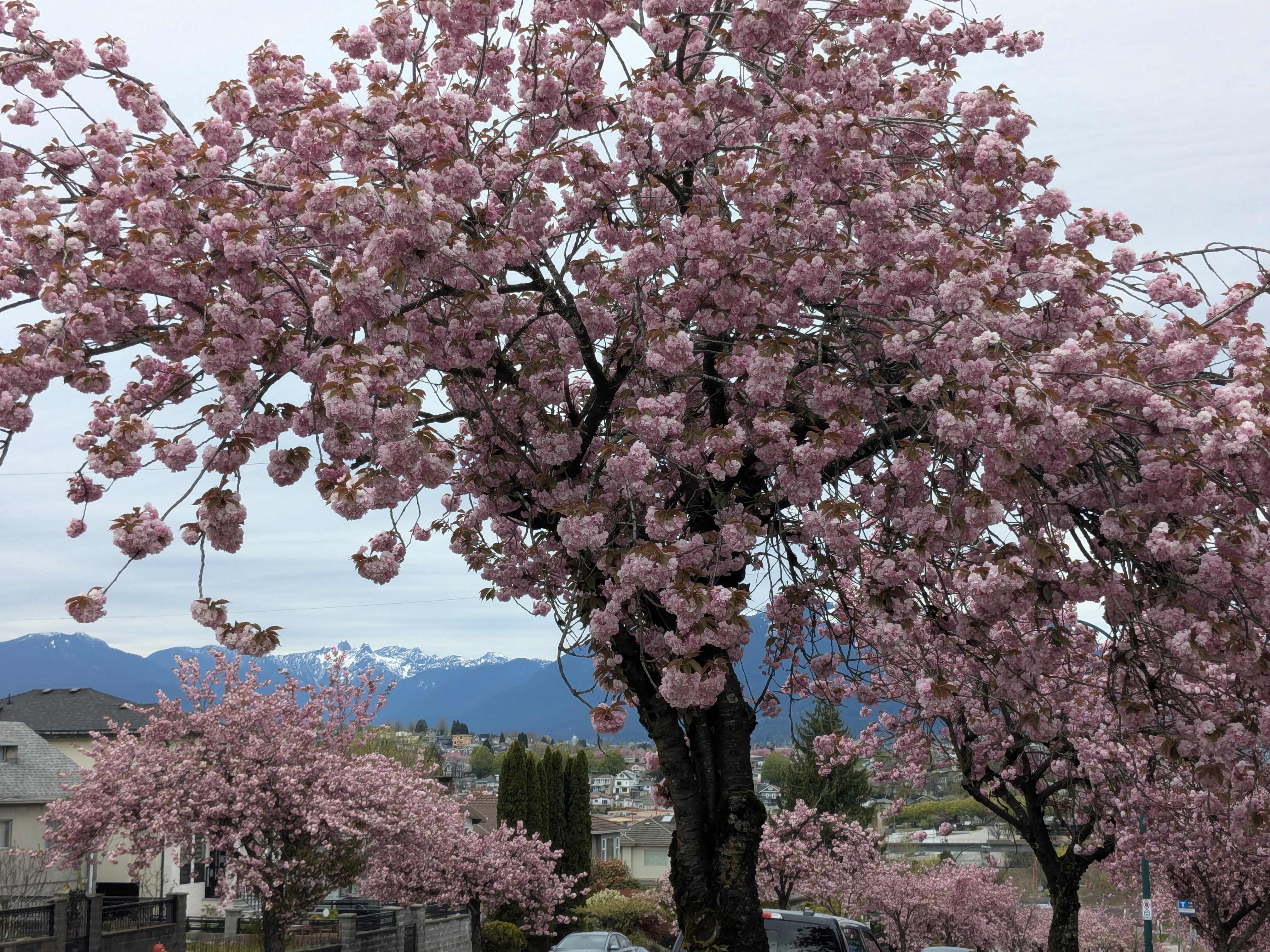

Results & Reflection
What We Did
- 5 open Vancouver datasets → ~3,500 Japanese flowering cherry trees
- Snapped trees to 50k street segments → density per 100m
- Interactive maps: neighbourhood choropleth + top 30 streets
- TSP: 10-stop running route and 30-stop cycling tour
- GPX files ready to load into Strava / Garmin
Things Worth Remembering
st_nearest_feature()- point-to-line assignment in one call- Density without a minimum count is just noise from tiny alleys
- Neighbourhood and street analysis ask different questions - both matter
- TSP needs a cost matrix - Euclidean is fast, road network is honest
- Clean the network (
to_spatial_subdivision+to_spatial_smooth) before routing - OSRM turns stop order into actual road geometry
Where Claude Code Helped
Claude Code helped throughout:
- Map aesthetics - described what I wanted, it wrote the
{mapgl}layers - Google Maps URLs - split 30-stop tour into 3 legs of 10; shareable links
- OSRM fallback - street data had geometry issues; Claude found OSRM as an alternative
- Code comments - “why” not just “what”
- Porting analysis to slides - scripts → slide-friendly code chunks
- These slides - structure, submodule wiring, Kyle Walker shout-out
- ASCII diagrams - mock up flow and structure diagrams
- Can also use Quarto + Mermaid diagrams
The code is not the hard part
AI can write the code. What it cannot do: ask the right question, recognize a wrong result, or know which assumption to challenge next. That is the skill worth building.
Open Data is Amazing
Vancouver Open Data Portal - free, well-documented, kept up to date:
- Tree locations and dimensions
- Street network (block-level segments)
- Neighbourhood boundaries
- Hundreds more datasets
Sharing Results with Quarto
Same R code → blog post via Quarto + GitHub Pages
- Interactive maps embedded directly
- GPX download links
- No R required to read it
- Analysis repo, talk, and blog are all Quarto documents
The Quarto stack
analysis.R
↓
results/README.qmd ← blog post source
↓ quarto render
docs/index.html ← GitHub PagesGrace’s talk from last month covers exactly this.
Appreciating City Planners
The city has been deliberate about where these trees go:
- Spread across many neighbourhoods, not concentrated in one spot
- Different species bloom at different times → longer season overall
- Even a single tree at an intersection makes a difference
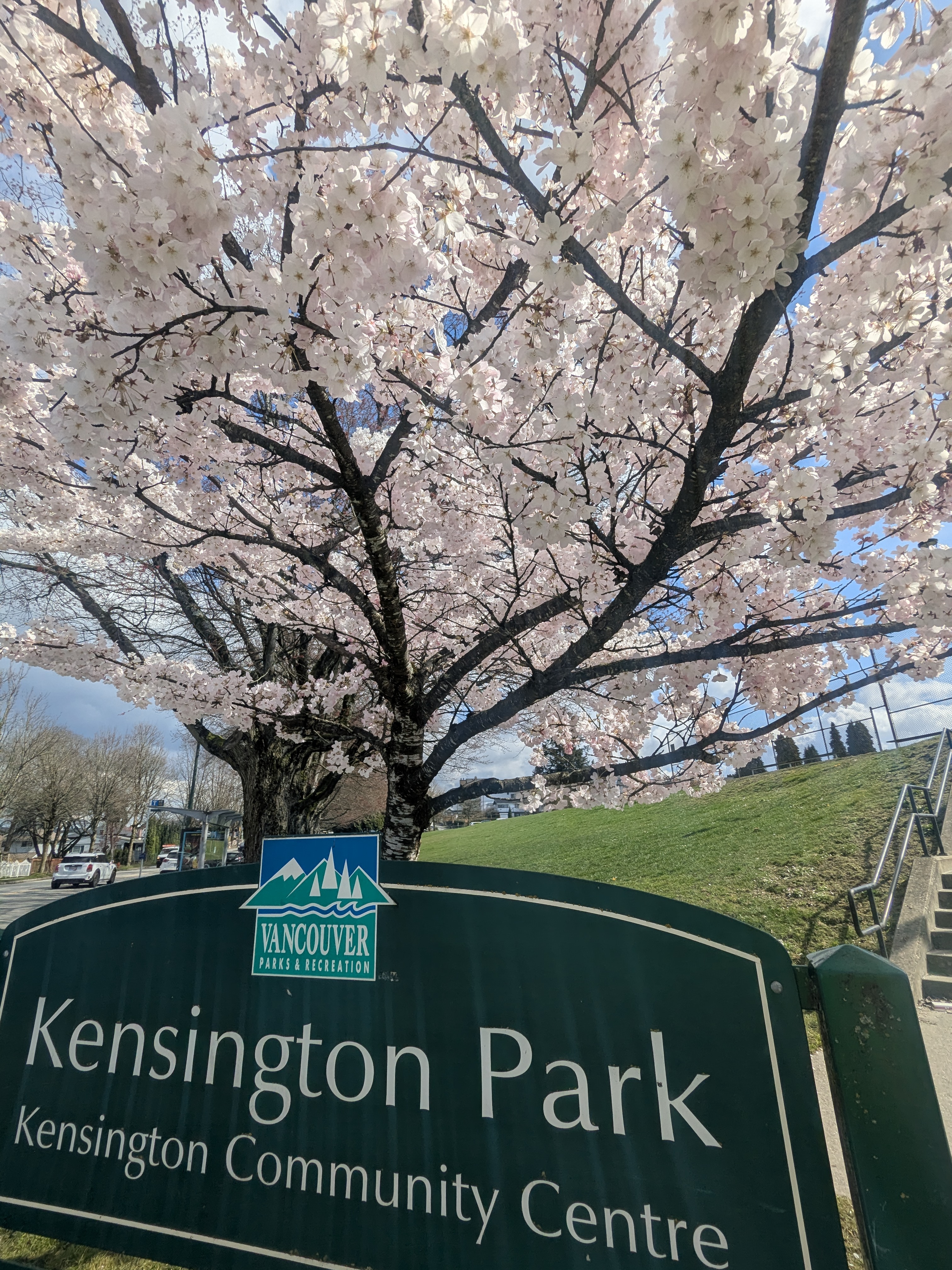

Next: Bloom-Aware Routes
The problem: routes show where trees are planted, not where they’re blooming
Confirmed on my March 30th run.
The idea
- Crowd-sourced bloom photos: Facebook, Reddit, vcbf.ca
- Estimate bloom timing per species
- Species matters as much as microclimate
- Build routes for trees actually blooming on a given day
What this becomes
Pick a date → route of currently-blooming streets:
- Early March → early-blooming species
- Peak season → full grand tour
- Late April → late-blooming varieties only
The Blossoms Don’t Last Long
This season is over!
Resources
- Analysis code: github.com/chendaniely/yvr-cherry-blossoms
- Blog post: chendaniely.github.io
- GPX files in the repo
data/routes/
R-Ladies YVR · https://github.com/chendaniely/rladies-2026-04-30-blossoms